Nanobodies – Their Functional Mechanisms and Engineering Novel Nanobodies
“Assoc. Prof. Mert Gur of the Mechanical Engineering Department utilizes all-atom molecular dynamics simulations to investigate the effectiveness of nanobodies, which are single domain antibodies, against SARS-CoV-2 variants, determines how their effectivity can be increased, and designs nanobodies specific to the variants and tests them via computer simulations. “
Nanobodies are single domain antibodies that can bind their drug targets strongly as regular antibodies, easily enter the cell due to their small size, and be mass-produced at higher rates with lower costs than antibodies. Their stability resembles larger sized antibodies. Therefore, using nanobodies in diagnostics and therapeutics is expected to have an important place in future of medicine and health sciences.
Caplacizumab, the first therapeutic nanobody, was recently (2019) approved by the US Food and Drug Administration. A growth is expected in development and use of nanobody-based drugs and diagnostic kits. Recent studies showed that nanobodies can bind SARS-CoV-2 Spike glycoprotein and prevent SARS-CoV-2 from infecting cell. Since here is a scarcity in drugs preventing SARS-CoV-2 infection, this success of nanobodies in preventing cell infection made great impact on the field.
Nanobodies differ mainly in their complementarity determining regions (CDRs), which are loop regions that have exquisite chemical composition and shape that are complementary to their protein targets. CDRs comprise only 20-30% of the nanobody structure and the remaining core is mostly conserved. Thus, nanobodies can be engineered by programming their CDR sequences, likely without significantly affecting their stability. This property of nanobodies would be useful in fighting diseases and epidemics such as SARS-CoV-2 where drug targets undergo mutations continuously.
Although many countries such as US, Germany and Turkey have reached high vaccination ratios, COVID-19 cases still keep increasing due to the resistance of the SARS-CoV-2 variants (currently Delta and Omicron) to all available vaccines. All available vaccines promote antibody production against the SARS-CoV-2 Spike glycoprotein, and most vaccine induced antibodies showed reduced or completely lost neutralization activity against the spike protein of the Delta variant. However, the effectiveness of neutralizing nanobodies, which are single-domain antibodies, against Delta Spike remains to be explored. Gur and his colleagues have been investigating the binding strengths of available nanobodies to the spike protein of SARS-CoV-2 variants and their ability to disrupt ACE2 binding of the spike protein. Their simulations predicted a clear decrease in binding strength and inhibition effectivity for these nanobodies.
New variants are expected to emerge due to the natural course of the pandemic, posing the danger of reducing nanobody and antibody binding efficiency further to critically low levels. Therefore, there is a need to design novel nanobodies specific to current and possible future SARS-CoV-2 variants. Gur and his colleagues are testing the effectiveness of nanobodies, known to be effective against SARS-CoV-2, against the SARS-CoV-2 variant using molecular dynamics simulations. In addition, by applying protein engineering, novel nanobodies that bind effectively to delta and omicron variants are designed and tested with molecular dynamics simulations.
Gur has been acting as the co-PI of two COVID-19 High Performance Computing Consortium projects performed in collaboration with UC Berkeley (USA) with the aim of exploring the effectiveness of nanobodies to in preventing ACE2 interactions with the Spike protein of SARS-CoV-2 and its variants, and to also design novel nanobodies specific to SARS-CoV-2 variants.
Gur and his colleagues investigated binding between the SARS-CoV-2 Alpha, Beta and Delta variant Spike protein and nanobodies H11-H4, H11-D4 and Ty1 by running more than 25 µs of molecular dynamics simulations (~20 million core hours). These simulations predicted a clear decrease in binding strength for these nanobodies. They also provided designed parameters to improve nanobody binding to the Delta Spike protein. Gur and his colleagues are currently investigating how mutations on the Delta and Omicron Spike variant alter Spike-nanobody interactions and how these nanobodies can be reengineered to increase their effectiveness against Delta Spike. Gur and his colleagues are engineering and computationally testing novel nanobodies against SARS-CoV-2 delta and omicron variants. He has performed some of the first molecular dynamics simulations of nanobody-spike protein interactions and performed a significant contribution to the field since nanobody interactions with WT and variant spike proteins have not been simulated at atomistic detail before.
The size of the simulated systems comprising the spike protein, ACE2 and nanobody are at least ~10-100 times larger than the system sizes in most molecular dynamics simulations performed in the literature. Considering that a very high number of nanobodies have to be tested and the number of variants is constantly increasing, only a limited number of research groups have the massive resources to perform this work. These resources are currently provided to Gur by the COVID-19 High Performance Computing Consortium.
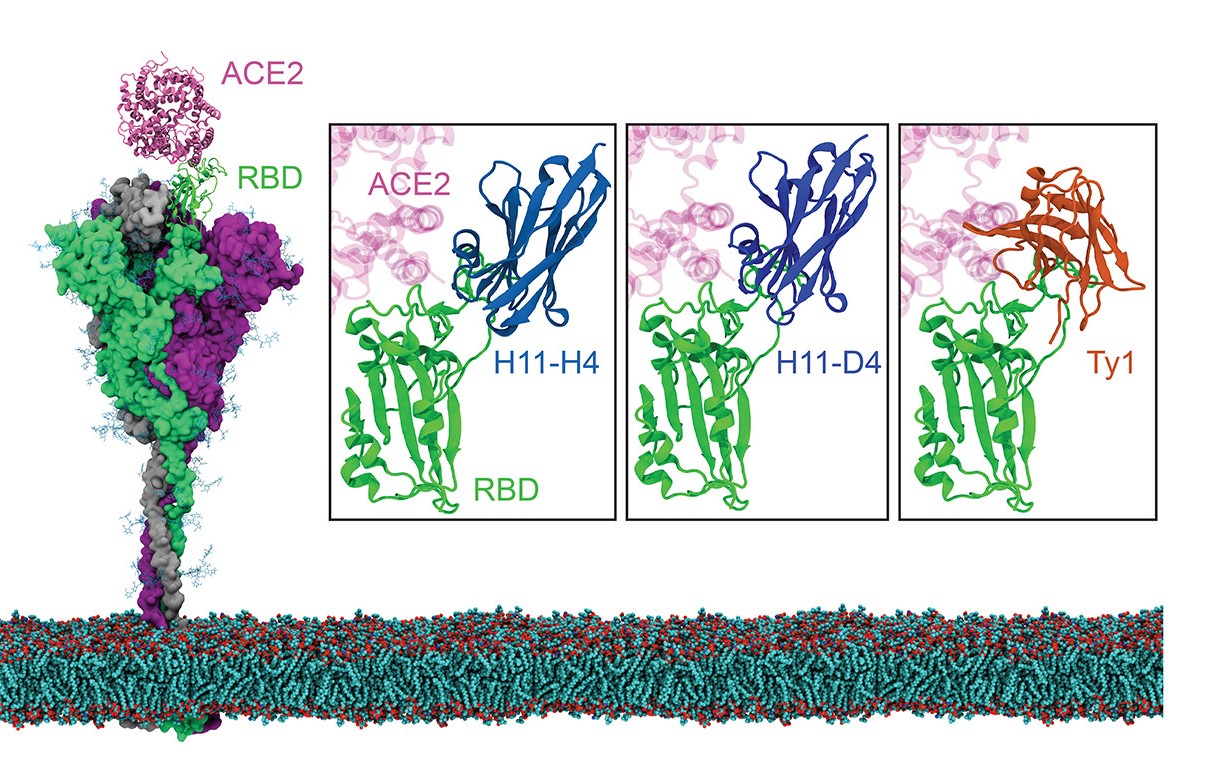
Figure 1. Nanobodies in complex with the RBD of the S protein located on the viral membrane. The atomic models were constructed using the full-length S protein model and the structure of RBD in complex with ACE2 (PDB 6M0J). The S protein trimer shown is composed of two protomers with RBD in the down position (gray and purple) and one protomer with RBD in the up position (green). The structures of RBD in complex with the nanobodies H11-H4, H11-D4, and Ty1 (PDB 6ZBP, 6YZ5, and 6ZXN, respectively) are highlighted in boxes.

Video 1. The electrostatic repulsion between ACE2 and H11-H4 upon H11-H4 docking and ACE2 discolation for WT and variants.
Related Articles:
1.Golcuk M , Yildiz A., Gur. M. (2022). Omicron BA.1 and BA.2 Variants Increase the Interactions of SARS-CoV-2 Spike Glycoprotein with ACE2. The Journal of Molecular Graphics & Modelling. 108286.
2. Golcuk M., Hacisuleyman, A., Yilmaz, S. Z., Taka, E. Yildiz A., Gur. M. (2022). SARS-Cov-2 Delta Variant Decreases Nanobody Binding and ACE2 Blocking Effectivity. Journal of Chemical Information and Modeling. 62 (10), 2490-2498.
3. Golcuk, M., Hacisuleyman, A., Erman, B., Yildiz, A., Gur, M. (2021). Binding mechanism of neutralizing Nanobodies targeting SARS-CoV-2 Spike Glycoprotein. Journal of Chemical Information and Modeling. 61(10), 5152–5160.
Fundings:
UHeM, COVID-19 HPC Consortium (Grant No. TG-MCB200070, TG-BIO210181)
Collaboration:
Ahmet Yıldız (UC Berkeley)
Lab Members Involved:
Mert Gölcük, Sema Zeynep Yılmaz, Clara Xazal Buran, Reyhan Metin Akkaya