Protein–Protein Interactions and Drug Design
Protein-protein interactions play a key role in the function of every living organisms. Novel discoveries in molecular disease pathways have led to an increasing interest in the design of drugs to modulate protein-protein interactions along those molecular disease pathways; as opposed to those only targeting the active site within a single enzyme/protein. Our lab utilizes computational modeling methods as well as statistical mechanics to model protein-protein interactions mechanisms and change of protein structure/dynamics/function upon ligand/protein binding. Critical insight/information on the protein-protein interaction mechanisms and its effects are elucidated that is required for the development/design of drugs to disrupt and inhibit those interactions. In silico drug design is performed to this aim. Computational simulations to understand protein-protein interactions and drug/peptide design are performed both at the molecular level, including Molecular Dynamics and Docking simulations, and at coarse grained level by utilizing methods such as GNM and ANM.

Related Articles:
1. Gur, M., Taka, E., Yilmaz, S. Z., Kilinc, C., Aktas, U., Golcuk, M. (2020). Conformational transition of SARS-CoV-2 spike glycoprotein between its closed and open states. The Journal of Chemical Physics, 153(7), 075101.
2. Gur, M., Golcuk, M., Gul, A., Erman, B. (2020). Molecular dynamics simulations provide molecular insights into the role of HLA-B51 in Behçet’s Disease pathogenesis. Chemical Biology & Drug Design, 96(1), 644-658.
3. Zhou, Z., Feng, Z., Hu, D., Yang, P., Gur, M., Bahar, I., Cristofanilli, M., Gradishar, W.J., Xie, X., Wan, Y. (2019). A novel small-molecule antagonizes PRMT5-mediated KLF4 methylation for target therapy. EBioMedicine-The Lancet, 44, 98-111.
4. Gur, M., Blackburn, E. A., Ning, J., Narayan, V., Ball, K. L., Walkinshaw, M. D., Erman, B. (2018). Molecular dynamics simulations of site point mutations in the TPR domain of cyclophilin 40 identify conformational states with distinct dynamic and enzymatic properties. The Journal of Chemical Physics, 148(14), 145101.
5. Hu, D., Gur, M., Zhou, Z., Gamper, A., Hung, M. C., Fujita, N., Lan L , Bahar I., Wan, Y. (2015). Interplay between arginine methylation and ubiquitylation regulates KLF4-mediated genome stability and carcinogenesis. Nature Communications, 6, 8419. (Joint corresponding author)
6. Zomot, E., Gur, M., Bahar, I. (2015). Microseconds simulations reveal a new sodium-binding site and the mechanism of sodium-coupled substrate uptake by LeuT. Journal of Biological Chemistry, 290(1), 544-555.
7. Das, A., Gur, M., Cheng, M. H., Jo, S., Bahar, I., Roux, B. (2014). Exploring the conformational transitions of biomolecular systems using a simple two-state anisotropic network model. PLOS Computational Biology, 10(4), e1003521.
8. Gur, M., Madura, J. D., Bahar, I. (2013). Global transitions of proteins explored by a multiscale hybrid methodology: application to adenylate kinase. Biophysical Journal, 105(7), 1643-1652.
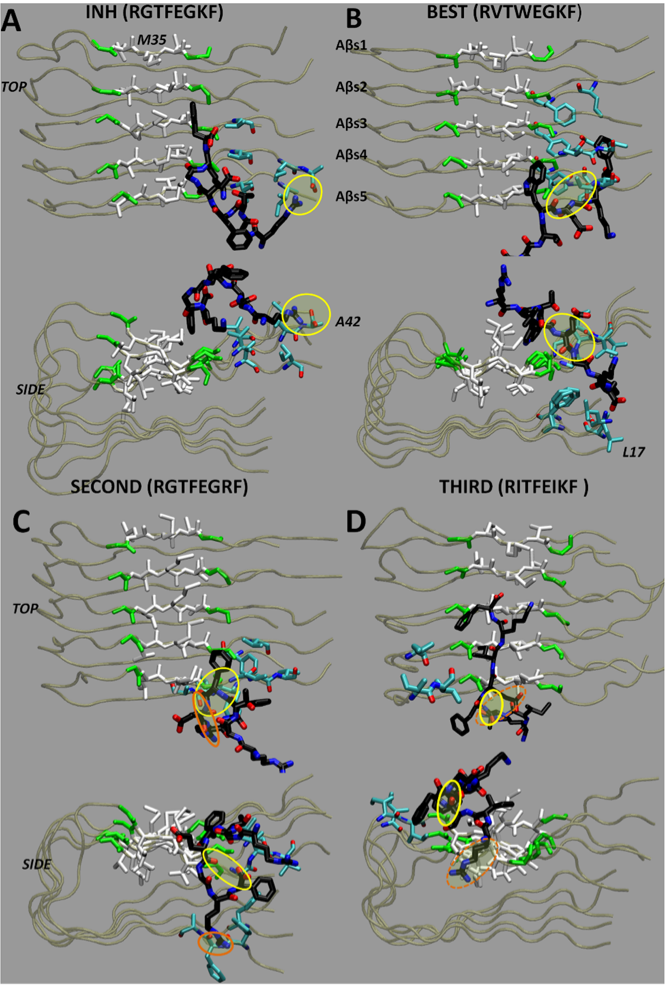
9. Eskici, G., Gur, M. (2013). Computational design of new peptide inhibitors for amyloid beta (Aβ) aggregation in Alzheimer's disease: application of a novel methodology. PLOS One, 8(6), e66178.
10. Meireles, L., Gur, M., Bakan, A., Bahar, I. (2011). Pre‐existing soft modes of motion uniquely defined by native contact topology facilitate ligand binding to proteins. Protein Science, 20(10), 1645-1658. (Joint corresponding author)
Lab members involved:
Mert Golcuk, Sema Zeynep Yilmaz, Reyhan Metin Akkaya, Clara Xazal Buran, Derman Basturk, Ebru Tuncay, Ayla Eren, Cihan Ugur Otcu